A new synthesis finds that common epilepsies are driven by thousands of tiny-effect genetic variants, most still undetected.
A modestly larger whole genome association study (GWAS) for genetic generalized epilepsy could capture approximately 50% of its common genetic variance, making it remarkably cost-efficient.”
OSLO, ØSTLANDET, NORWAY, March 10, 2026 /EINPresswire.com/ -- An insightful mini-review published in Genomic Psychiatry synthesizes the rapidly expanding landscape of molecular genetic research on common epilepsies, assembling evidence from genome-wide association studies (large-scale genetic scans across the human genome), whole-exome sequencing projects, and advanced statistical modeling to illuminate the polygenic architecture that underpins these heterogeneous neurological disorders. The synthesis, led by Dr. Olav B. Smeland of the Centre for Precision Psychiatry at Oslo University Hospital and the University of Oslo, draws a detailed portrait of a genetic landscape far more intricate than early twin studies ever suggested.— Dr. Olav B. Smeland, Center for Precision Psychiatry, University of Oslo
Epilepsy is not one disease. It is a constellation of seizure disorders that affects approximately 50 million people globally and carries increased mortality, psychiatric comorbidity, and, for roughly one-third of patients, resistance to existing medications. What determines who develops epilepsy, which subtype emerges, and who responds to treatment? Those questions have driven genetic research along two parallel tracks for decades. One track has yielded dramatic results: studies of severe monogenic epilepsies, such as developmental and epileptic encephalopathies, have identified over a thousand implicated genes. The other track, focused on common epilepsies including genetic generalized epilepsy and focal epilepsy, has moved more slowly, hindered by the sheer complexity of polygenic inheritance.
Twin studies set the stage for modern epilepsy genetics. The review traces the evidentiary thread back to the 1930s, when researchers first demonstrated higher concordance rates for epilepsy among identical twins compared with fraternal twins. The largest such study, encompassing 47,626 twin pairs, found concordance of 28% in identical twins versus 7% in fraternal twins. The numbers diverge further when epilepsy subtypes are examined separately. For genetic generalized epilepsy, identical-twin concordance reached 77% and fraternal-twin concordance was 35%; for focal epilepsy, those figures dropped to 40% and 3%, respectively. Modern molecular methods have quantified this heritability in a different currency: the fraction of phenotypic variation attributable to common genetic variants is estimated to be approximately three times larger for genetic generalized epilepsy than for focal epilepsy. Specific subtypes such as juvenile myoclonic epilepsy and childhood absence epilepsy show even higher heritability estimates, underscoring that diagnostic precision matters enormously in genetic research.
Rare genetic variants also contribute a convergent but distinct story. The review describes how rare variants, those with a minor allele frequency below 1%, also contribute to epilepsy risk, although they are present in only a minority of cases. A study involving 13,420 epilepsy cases demonstrated increased copy number variant burden across all common epilepsy types compared with controls, with genetic generalized epilepsy showing the highest burden. Recurrent deletions at the 15q13.3 chromosomal locus emerged as the strongest risk factor for genetic generalized epilepsy, with an odds ratio of 36.04. Whole-exome sequencing studies have identified protein-truncating ultrarare variants in genes encoding components of the GATOR1 complex, a negative regulator of the mTORC1 cellular growth-signaling pathway, as robust contributors to non-acquired focal epilepsy risk. What makes these findings especially compelling is the convergence between rare and common variant signals. Genes like DEPDC5, NPRL3, SCN1A, and SCN8A appear in both rare variant analyses and common variant association studies, pointing toward shared biological pathways involving ion channel function, synaptic excitability, and excitatory-inhibitory balance.
The common variant landscape has expanded substantially in recent years. The largest genome-wide association study of common epilepsies to date, conducted by the International League Against Epilepsy with 29,944 cases and 52,538 controls, identified 26 genome-wide significant loci. The distribution was strikingly uneven: 22 loci were associated with genetic generalized epilepsy from only 7,407 cases, while focal epilepsy, despite more than twice as many cases at 16,384, yielded no genome-wide significant associations. This asymmetry is not merely a matter of sample size, the review authors argue, but reflects fundamental differences in genetic architecture between subtypes. Among the 29 potential causal genes prioritized from those 26 loci, ten are established monogenic epilepsy genes, including ion channel subunits such as SCN8A, SCN1A, CACNA1I, and KCNN2, along with neurotransmitter receptor components GABRA2 and GRIK1. This genetic convergence between monogenic and polygenic forms of epilepsy is perhaps the review's most striking integrative insight.
Epilepsy and psychiatry converge at the genomic level in ways that challenge traditional diagnostic boundaries. The review devotes substantial attention to genetic pleiotropy, the phenomenon whereby genetic variants influence more than one trait or disorder simultaneously. The genome-wide genetic correlation between focal epilepsy and genetic generalized epilepsy is 0.61, indicating that many common variants jointly increase risk for both. Both subtypes also show moderate negative genetic correlations with cognitive ability, consistent with the well-documented cognitive impairment seen in epilepsy patients. Using the bivariate MiXeR model, a statistical framework for quantifying genetic overlap, the review authors demonstrate that most variants associated with genetic generalized epilepsy are also associated with major psychiatric disorders, including schizophrenia, major depression, bipolar disorder, and anxiety.
"The extensive genetic overlap between epilepsy and psychiatric disorders provides a molecular explanation for what clinicians have long observed at the bedside," said Naz Karadag, first author and researcher at the Department of Molecular Medicine, Institute of Basic Medical Sciences, University of Oslo. "Understanding these shared genetic foundations may eventually help identify epilepsy patients at elevated risk for psychiatric comorbidities."
The review also highlights a less intuitive finding: approximately 30% to 40% of the common variants contributing to epilepsy risk overlap with variants influencing cortical thickness and cortical surface area, despite the absence of significant genome-wide genetic correlations between these phenotypes. The relationship is more tangled than simple directional associations can capture.
On the clinical front, the review is carefully measured. Genetic testing is currently established for severe early-onset or syndromic epilepsies, where identifying a pathogenic variant may guide treatment. But for common epilepsies, with their complex inheritance and the fact that only a minority of cases harbor rare pathogenic variants of large effect, routine genetic testing remains premature. Polygenic risk scores, which estimate an individual's genetic susceptibility based on thousands of common DNA variants, offer a different angle. The lifetime risk of epilepsy increases by a hazard ratio of 1.73 per standard deviation increase in a genetic generalized epilepsy polygenic risk score, a figure comparable to polygenic risk prediction in cardiology. Yet the discriminative performance remains insufficient for population screening.
"Polygenic risk scores for epilepsy show promise in specific clinical contexts, such as risk stratification after a first unprovoked seizure," said Dr. Smeland. "But we must be cautious. Current scores should not be used for routine clinical decision-making, and broadening ancestral diversity in our study populations is essential before any implementation can be considered equitable."
A critical equity gap persists: over 92% of cases in the largest epilepsy genome-wide association study are of European ancestry, severely limiting the generalizability of risk scores across populations.
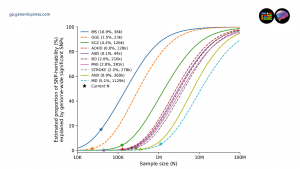
One of the review's most compelling analytical contributions involves power projections generated using the MiXeR framework. At current sample sizes for genetic generalized epilepsy (effective N of approximately 23,000), only about 1.5% of SNP-heritability is explained by genome-wide significant variants. If genetic generalized epilepsy genome-wide association studies were scaled to comparable sample sizes used in stroke research (an effective N of approximately 278,000), roughly 50% of common genetic variance could be captured. The investment required is smaller than for most other complex brain disorders, and the yield would be disproportionately large.
The review does not shy away from limitations. Existing datasets are predominantly European and drawn from a narrow range of sources. The role of somatic mosaicism in common epilepsies remains largely unexplored. The statistical power for focal epilepsy genome-wide association studies is currently insufficient for reliable MiXeR analysis, leaving a significant portion of the epilepsy landscape unmapped. And the phenotypic categories used in large-scale genetic studies may not capture the clinical granularity needed to detect subtype-specific genetic signals.
"We are still at an early stage of genetic discovery for common epilepsies," said Julian Fuhrer, co-author and researcher at the Centre for Precision Psychiatry, Oslo University Hospital and University of Oslo, who generated all data analyses and figures for the review. "The genetic signal is there, the tools are improving, and the returns on investment in larger, more diverse samples are clear. What we need now is the coordinated effort to make it happen."
Looking forward, the review envisions a future in which genetics is integrated with other data modalities, including clinical and cognitive variables, other omics data, electronic health records, neuroimaging, electrophysiology, and sensing device phenotypes, to construct genuinely multimodal prediction models. Large biobanks with longitudinal data, such as the UK Biobank and the All of Us Research program, will be essential platforms. Rapidly advancing artificial intelligence and machine-learning algorithms may provide the computational means to integrate these diverse data streams effectively.
The synthesis by Dr. Smeland and colleagues, including co-authors Dr. Kjell Heuser of the Department of Neurology at Oslo University Hospital and Professor Ole A. Andreassen of the Centre for Precision Psychiatry and the KG Jebsen Centre for Neurodevelopmental Disorders at the University of Oslo, represents a clear-eyed inventory of what the field knows, what it suspects, and what it still cannot answer.
The peer-reviewed mini-review in Genomic Psychiatry titled "The genetics of common epilepsies" is freely available via Open Access, starting on 10 March 2026 in Genomic Psychiatry at the following link: https://doi.org/10.61373/gp026y.0027
The full reference for citation purposes is: Karadag N, Fuhrer J, Heuser K, Andreassen OA, and Smeland OB. The genetics of common epilepsies. Genomic Psychiatry 2026. DOI: https://doi.org/10.61373/gp026y.0027. Epub 2026 Mar 10.
About Genomic Psychiatry: Genomic Psychiatry: Advancing Science from Genes to Society (ISSN: 2997-2388, online and 2997-254X, print) represents a paradigm shift in genetics journals by interweaving advances in genomics and genetics with progress in all other areas of contemporary psychiatry. Genomic Psychiatry publishes peer-reviewed medical research articles of the highest quality from any area within the continuum that goes from genes and molecules to neuroscience, clinical psychiatry, and public health.
Visit the Genomic Press Virtual Library: https://issues.genomicpress.com/bookcase/gtvov/
Our media website is at: https://media.genomicpress.com/
Our full website is at: https://genomicpress.com/
Ma-Li Wong
Genomic Press
mali.wong@genomicpress.com
Visit us on social media:
X
LinkedIn
Bluesky
Instagram
Facebook
Legal Disclaimer:
EIN Presswire provides this news content "as is" without warranty of any kind. We do not accept any responsibility or liability for the accuracy, content, images, videos, licenses, completeness, legality, or reliability of the information contained in this article. If you have any complaints or copyright issues related to this article, kindly contact the author above.